“Seeing” the toxic huntingtin protein in people with HD
New tools let us “see” clumps of toxic huntingtin protein which build up in the brains of people with Huntington’s disease over time. Tracking these clumps might help us to better understand how HD progresses or how treatments might slow or halt HD.

Scientists have developed a tool which allows us to “see” the toxic clumps of the huntingtin protein using special scanners. People with Huntington’s disease (HD) make a toxic form of the huntingtin protein which forms clumps in cells of their bodies, which accumulate during HD progression. Tracking how these clumps form over time in people with HD, or how they change when people with HD take different treatments, could help us better understand the progression of HD and which medicines help patients most.
What are these protein clumps?
We all have 2 copies of the huntingtin gene but for people who have Huntington’s disease (HD), one of their copies has a type of mutation called a repeat expansion. This mutation occurs in a repetitive bit of the huntingtin gene DNA code which has the letters “C”, “A” and “G” repeating over and over. If you don’t have HD, you will have less than ~35 CAG repeats in your huntingtin gene but for people with HD, the mutation means they will have more than 35 CAG repeats in one of their huntingtin genes.
The huntingtin gene is the recipe our cells use to make the huntingtin protein so if the DNA code of this recipe is changed, the protein which our bodies make will also be changed. Proteins are made from long strings of chemicals called amino acids, following the instructions laid down in our DNA. The DNA letters “CAG” are the recipe notes for the amino acid glutamine. This means that if the CAG repeat number gets bigger, the huntingtin protein will have many more repeating glutamines. Huntingtin protein molecules with too many glutamines can’t assemble properly so can form toxic clumps.
We have known about these clumps for a long time now and they can be seen in the brains of people with HD when we look under a microscope. However, tracking these clumps in living patients has been challenging and most of our knowledge of them comes from looking at them in post-mortem brain samples from animal models of HD or patients who have graciously donated their brains to research.
Why do we want to look at these pesky clumps?
Scientists from many labs across the UK, Germany, Italy, Sweden, and the US have developed molecular tools which now allow us to “see” these clumps in living animals, and hopefully soon, HD patients. These tools bind to the huntingtin protein clumps and have chemical decorations, called radiolabels, which mean that they light up when looked at by PET (positron emission tomography) scan.
This type of molecular tools are known as PET tracers and are used in lots of different medical and diagnostic settings to allow doctors and researchers to image specific parts of your body. Different types of tracers can be swallowed, injected, or inhaled depending on what part of your body is being looked at. Once the patient has the PET tracer in their body, they will be scanned, and the part of the body being looked at will light up if the target of the PET tracer is there because the tracer is slightly radioactive. Similar tools have been developed for studying other diseases such as Pittsburgh compound B which is used to look at similar clumps in people with Alzheimer’s.
Making PET tracers which allow researchers to see the toxic clumps of huntingtin protein is an attractive idea for several reasons. Firstly, a PET scan can be performed on patients at multiple time points throughout their life so we can track how the clumps accumulate over time throughout the progression of HD. Many of our current methods for looking at huntingtin clumps in patient’s brain can only currently be done at the very end of the disease on post-mortem tissue.
Secondly, PET scans are non-invasive procedures and allow us to look right in the brain whereas more intrusive procedures like measuring huntingtin protein in spinal fluid provide only a proxy for what we think is happening in the brain. Thirdly, the clumps are formed from the toxic form of the huntingtin protein so PET scans will allow researchers to specifically measure changes to this specific form of mutant huntingtin. This differs from most of ways we measure and analyse huntingtin in spinal fluid or blood which measure all the different forms of huntingtin, including the healthy huntingtin protein.
Development of the first huntingtin PET ligand
Last August, an early version of these tools was published called CHDI-180R – the first time a PET tracer has been made for the huntingtin protein! A team led by Celia Dominguez at CHDI Foundation showed that the tool molecule CHDI-180R was able to bind very tightly to the toxic huntingtin protein clumps in a test tube. They also used CHDI-180R to show where the clumps of toxic huntingtin were in brain samples from HD mouse models.
In the brains of mice with the HD mutation, clumps of the toxic huntingtin protein could be seen in many different brain regions which are known to be affected by HD, whereas in mice without the HD mutation, these clumps could not be seen, even though they had also been injected with the tool molecule CHDI-180R. Finally, the scientists showed that CHDI-180R spread well through the brain and was also safe with no side effects in both monkeys and rats.
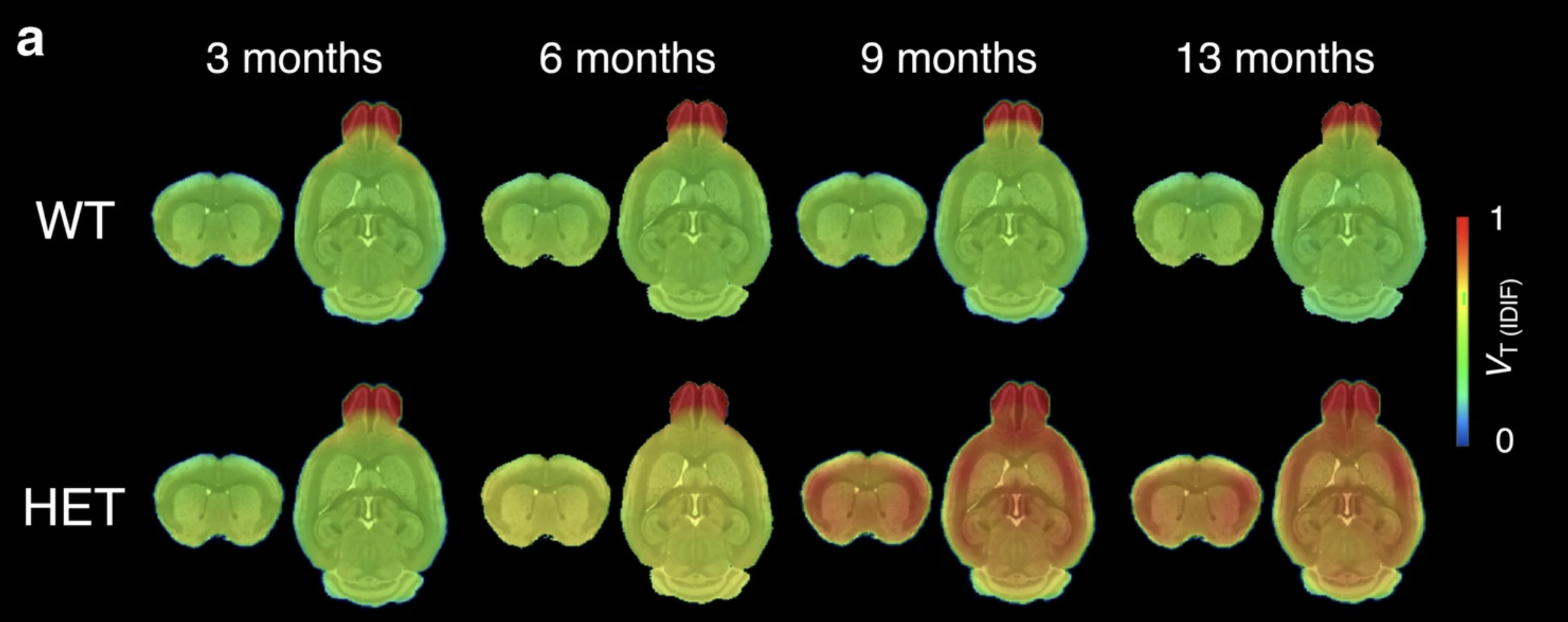
Image credit: Bertoglio et al (2021) Eur J Nucl Med Mol Imaging
Fine-tuning the tools
PET tracer development often takes multiple attempts before an optimal tool is found so the same international group of scientists is also developing other versions of this tracer to have lots of back up options. These (hopefully) new and improved versions of the molecular tool are being tested to work out how they spread in the brains of animals tested.
Other diseases like Alzheimer’s also have protein clumps which build up in nerve cells, but these are made up of other potentially toxic proteins, like amyloid beta. The scientists are also checking how specific these tools are for the huntingtin protein clumps which accumulate over time in HD patients compared to other disease protein clumps, like those from Alzheimer’s patients. So far, the results have been very encouraging, so the scientists are now keen to start testing the tracers in people.
So, what’s next?
A clinical trial is being conducted called iMagemHTT study, which will investigate the huntingtin tracer in people. The trial will use PET/MRI imaging to understand how the PET ligand tracks huntingtin in the brain. We previously reported on some encouraging preliminary data from the Phase I study of this tracer at the CHDI virtual meeting earlier this year. So far, the findings are encouraging, so they are continuing to add participants to the study.
The amount of huntingtin clumps in the brains of people with HD is a good biomarker of disease progression. Biomarkers are objective measurements scientists and clinicians can take to track HD’s progression which can be important for working out the best treatment options, as well as if treatments are working properly. It is possible that HD patients in the future might be monitored by PET scan using these type of tools.
If the PET ligands work as scientists hope, it could also be used to track huntingtin-lowering in the brain in future trials. Despite some setbacks, huntingtin lowering is still a promising strategy for treating HD which is being pursued by Novartis, PTC Therapeutics, Wave and Uniqure, all of whom have clinical trials underway. Regardless of what happens with Huntingtin lowering, these exciting new tools are giving scientists the ability – for the first time ever – to track mutant Huntingtin protein across the entire brain of living HD patients, which is a huge advance.
We look forward to updating you more on this topic soon!
Learn more
Sources & References
For more information about our disclosure policy see our FAQ…